Using single-cell genomics to study early development
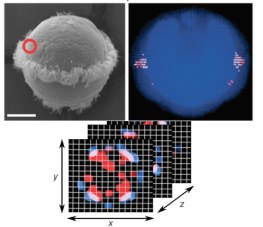
Gastrulation and the specification of the three germ layers are key events in animal development. However, molecular analyses of these processes have been limited due to the small number of cells present in gastrulating embryos. With recent developments in the field of single-cell biology however, it is now possible to overcome these limitations and to characterize, for the first time at the single-cell level, how cell fate decisions are made.
In this presentation I will discuss data generated to study cell fate specification in mouse, as well as the computational strategies we have developed to model such data. I will then illustrate how these data can provide insight into germ layer specification and early erythropoiesis.
Ulysse Herbach (LBMC, ENS-Lyon)
Toward gene network inference using single-cell data
Ghislain Durif (LBBE, Lyon)
Count matrix factorization for single cell data analysis
Next Generation Sequencing (NGS) data are characterized by their high dimensionality. Analysing such data is a statistical challenge and requires the use of dimension reduction approaches. Compression and especially matrix factorization methods show particular abilities concerning data visualization or data interpretation, for instance to expose latent structure such as complex multi-correlation between genes. However NGS data like gene expression profiles are very specific, being count matrices (as integer and non negative) with particular patterns. For instance, single cell data are characterized by dropout events, resulting in an artificial amplification of the number of zeros (called zero-inflation). Indeed, a null value in read counts may refer to an absence of read or to a failure in the experiment due to the short amount of genetic material available in a single cell. The huge amount of zeros create artificial correlations between genes, and it becomes necessary to use specific statistical tools to avoid false interpretation.
In this talk I will present a compression methods, based on factor model and suitable for zero-inflated count data. It uses Bayesian framework with particular count distribution (Gamma-Poisson) and variational inference for scalability and efficient computations. This approach can be viewed as a generalization of Principal Components Analysis (PCA) for non-Gaussian data.
SeMoVi will take place at salle de conférence du CBP - IXXI (LR6-M8) 14h

